-Search query
-Search result
Showing 1 - 50 of 176 items for (author: huang & hy)

EMDB-36453: 
Structural basis of transcriptional activation by the OmpR/PhoB-family response regulator PmrA
Method: single particle / : Lou YC, Huang HY, Chen C, Wu KP

EMDB-41182: 
Cryo-EM map of the Unmodified nucleosome core particle in 100 mM KCl with local resolution values
Method: single particle / : Huang SK, Kay LE, Rubinstein JL
EMDB-41183: 
Cryo-EM map of the PARylated nucleosome core particle in 100 mM KCl with local resolution values
Method: single particle / : Huang SK, Kay LE, Rubinstein JL
EMDB-41184: 
Cryo-EM map of the Unmodified nucleosome core particle in 5 mM KCl with local resolution values
Method: single particle / : Huang SK, Kay LE, Rubinstein JL
EMDB-41178: 
Cryo-EM map of the PARylated nucleosome core particle in 5 mM KCl with local resolution values
Method: single particle / : Huang SK, Kay LE, Rubinstein JL

EMDB-35609: 
Cryo-EM structure of phosphoketolase from Bifidobacterium longum in octameric assembly
Method: single particle / : Chang CW, Tsai MD

EMDB-35610: 
Cryo-EM structure of phosphoketolase from Bifidobacterium longum in dimeric assembly
Method: single particle / : Chang CW, Tsai MD

EMDB-35611: 
Cryo-EM structure of cyanobacteria phosphoketolase complexed with AMPPNP in dimeric assembly
Method: single particle / : Chang CW, Tsai MD

EMDB-35612: 
Cryo-EM structure of cyanobacteria phosphoketolase complexed with AMPPNPin dodecameric assembly
Method: single particle / : Chang CW, Tsai MD

EMDB-35613: 
Cryo-EM structure of cyanobacteria phosphoketolase
Method: single particle / : Chang CW, Tsai MD

EMDB-35617: 
Cryo-EM structure of cyanobacteria phosphoketolase in dodecameric assembly
Method: single particle / : Chang CW, Tsai MD
EMDB-29301: 
Neurotensin receptor allosterism revealed in complex with a biased allosteric modulator
Method: single particle / : Krumm BE, Diberto JF, Olsen RHJ, Kang H, Slocum ST, Zhang S, Strachan RT, Fay JF, Roth BL

EMDB-29302: 
CryoEM structure of Go-coupled NTSR1 with a biased allosteric modulator
Method: single particle / : Krumm BE, DiBerto JF, Olsen RHJ, Kang H, Slocum ST, Zhang S, Strachan RT, Fay JF, Roth BL
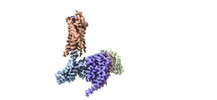
EMDB-29303: 
CryoEM structure of Go-coupled NTSR1
Method: single particle / : Krumm BE, DiBerto JF, Olsen RHJ, Kang H, Slocum ST, Zhang S, Strachan RT, Fay JF, Roth BL

PDB-8fmz: 
Neurotensin receptor allosterism revealed in complex with a biased allosteric modulator
Method: single particle / : Krumm BE, Diberto JF, Olsen RHJ, Kang H, Slocum ST, Zhang S, Strachan RT, Fay JF, Roth BL

PDB-8fn0: 
CryoEM structure of Go-coupled NTSR1 with a biased allosteric modulator
Method: single particle / : Krumm BE, DiBerto JF, Olsen RHJ, Kang H, Slocum ST, Zhang S, Strachan RT, Fay JF, Roth BL

PDB-8fn1: 
CryoEM structure of Go-coupled NTSR1
Method: single particle / : Krumm BE, DiBerto JF, Olsen RHJ, Kang H, Slocum ST, Zhang S, Strachan RT, Fay JF, Roth BL

EMDB-28092: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-093
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28090: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-040
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28091: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-045
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28093: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-156
Method: single particle / : Shek J, Callaway H, Li H, Yu X, Saphire EO

EMDB-28094: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-234
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28095: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-260
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28096: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-279
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28097: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-290
Method: single particle / : Yu X, Callaway H, Li H, Shek J, Saphire EO
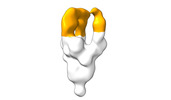
EMDB-28098: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-294
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28099: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-295
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28100: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-299
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28102: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-334
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
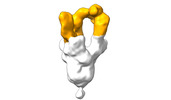
EMDB-28103: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-360
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28104: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-361
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28105: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-362
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28106: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-368
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28168: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-292
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28169: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-333
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
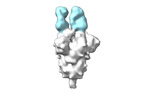
EMDB-28170: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-355
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28171: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-371
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
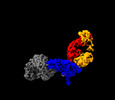
EMDB-28523: 
Structure of interleukin receptor common gamma chain (IL2Rgamma) in complex with two antibodies
Method: single particle / : Franklin MC, Romero Hernandez A

PDB-8epa: 
Structure of interleukin receptor common gamma chain (IL2Rgamma) in complex with two antibodies
Method: single particle / : Franklin MC, Romero Hernandez A

EMDB-32497: 
SARS-CoV-2 spike in complex with the ZB8 neutralizing antibody Fab (focused refinement on Fab-RBD)
Method: single particle / : Zeng JW

EMDB-32498: 
SARS-CoV-2 spike in complex with the ZB8 neutralizing antibody Fab (3U)
Method: single particle / : Zeng JW, Ge JW

EMDB-32499: 
SARS-CoV-2 spike in complex with the ZB8 neutralizing antibody Fab (2u1d)
Method: single particle / : Zeng JW, Wang XW

EMDB-16434: 
Type Ib beta-amyloid 42 Filaments from Human Brain
Method: helical / : Yang Y, Arseni D, Zhang W, Huang M, Lovestam SKA, Schweighauser M, Kotecha A, Murzin AG, Peak-Chew SY, Macdonald J, Lavenir I, Garringer HJ, Gelpi E, Newell KL, Kovacs GG, Vidal R, Ghetti B, Falcon B, Scheres HW, Goedert M

EMDB-26767: 
The 2.19-angstrom CryoEM structure of the [NiFe]-hydrogenase Huc from Mycobacterium smegmatis - Complex minus stalk
Method: single particle / : Grinter R, Venugopal H, Kropp A, Greening C
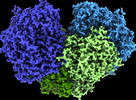
EMDB-26801: 
The 1.67 Angstrom CryoEM structure of the [NiFe]-hydrogenase Huc from Mycobacterium smegmatis - catalytic dimer (Huc2S2L)
Method: single particle / : Grinter R, Venugopal H, Kropp A, Greening C

EMDB-26802: 
The CryoEM structure of the [NiFe]-hydrogenase Huc from Mycobacterium smegmatis - Full complex focused refinement of stalk
Method: single particle / : Grinter R, Venugopal H, Kropp A, Greening C
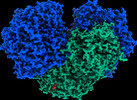
EMDB-27661: 
The 1.52 angstrom CryoEM structure of the [NiFe]-hydrogenase Huc from Mycobacterium smegmatis - catalytic dimer (Huc2S2L)
Method: single particle / : Grinter R, Venugopal H, Kropp A, Greening C

PDB-7utd: 
The 2.19-angstrom CryoEM structure of the [NiFe]-hydrogenase Huc from Mycobacterium smegmatis - Complex minus stalk
Method: single particle / : Grinter R, Venugopal H, Kropp A, Greening C

PDB-7uur: 
The 1.67 Angstrom CryoEM structure of the [NiFe]-hydrogenase Huc from Mycobacterium smegmatis - catalytic dimer (Huc2S2L)
Method: single particle / : Grinter R, Venugopal H, Kropp A, Greening C

PDB-7uus: 
The CryoEM structure of the [NiFe]-hydrogenase Huc from Mycobacterium smegmatis - Full complex focused refinement of stalk
Method: single particle / : Grinter R, Venugopal H, Kropp A, Greening C
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
